-Search query
-Search result
Showing 1 - 50 of 128 items for (author: tan & yz)

EMDB-36766: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36767: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36768: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36769: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36770: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36771: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36772: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36807: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36809: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D
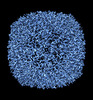
EMDB-36810: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36811: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36812: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36813: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D
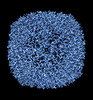
EMDB-36814: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36816: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36817: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36818: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36819: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36820: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36821: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36822: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-41230: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-41231: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-41695: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of RNA polymerase
Method: single particle / : Aiyer S, Shan Z, Oh J, Wang D, Tan YZ, Lyumkis D

EMDB-28540: 
BG505 UFO-E2p-L4P nanoparticle reconstructed by focused refinement with a mask around the nanoparticle core
Method: single particle / : Antanasijevic A, Zhang YN, Zhu J, Ward AB
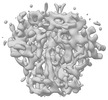
EMDB-28541: 
BG505 UFO trimer map reconstructed from BG505 UFO-E2p-L4P nanoparticle by localized reconstruction
Method: single particle / : Antanasijevic A, Zhang YN, Zhu J, Ward AB

EMDB-28542: 
BG505 UFO-10GS-I3-01v9-L7P nanoparticle reconstructed by focused refinement with a mask around the nanoparticle core
Method: single particle / : Antanasijevic A, Zhang YN, Zhu J, Ward AB

EMDB-28543: 
BG505 UFO trimer map reconstructed from BG505 UFO-10GS-I3-01v9-L7P nanoparticle by localized reconstruction
Method: single particle / : Antanasijevic A, Zhang YN, Zhu J, Ward AB

EMDB-33498: 
Cryo-EM structure of Sr35 resistosome induced by AvrSr35 R381A
Method: single particle / : Ouyang SY, Zhao YB, Li ZK, Liu MX

EMDB-33153: 
Cryo-EM structure of plant NLR Sr35 resistosome
Method: single particle / : Ouyang SY, Zhao YB, Li ZK, Liu MX

EMDB-33486: 
Cryo-EM structure of binary complex of plant NLR Sr35 and effector AvrSr35
Method: single particle / : Ouyang SY, Zhao YB, Li ZK, Liu MX

EMDB-26824: 
Citrus V-ATPase State 1, Highest-Resolution Class
Method: single particle / : Tan YZ, Rubinstein JL

EMDB-26825: 
Citrus V-ATPase State 1, H in contact with subunit a
Method: single particle / : Keon KA, Abdelaziz RA, Schulze WX, Schumacher K, Rubinstein JL

EMDB-26826: 
Citrus V-ATPase State 1, H in contact with subunits AB
Method: single particle / : Abdelaziz RA, Keon KA, Schulze WX, Schumacher K, Rubinstein JL

EMDB-26827: 
Citrus V-ATPase State 2, Highest-Resolution Class
Method: single particle / : Tan YZ, Rubinstein JL

EMDB-26828: 
Citrus V-ATPase State 2, H in contact with subunit a
Method: single particle / : Tan YZ, Rubinstein JL

EMDB-26829: 
Citrus V-ATPase State 2, H in contact with subunits AB
Method: single particle / : Tan YZ, Rubinstein JL

EMDB-26385: 
Structure of porcine V-ATPase with mEAK7 and SidK, Rotary state 2
Method: single particle / : Tan YZ

EMDB-26386: 
Structure of porcine kidney V-ATPase with SidK, Rotary State 1
Method: single particle / : Tan YZ

EMDB-26387: 
Structure of porcine kidney V-ATPase with SidK, Rotary State 2
Method: single particle / : Tan YZ

EMDB-26388: 
Structure of porcine kidney V-ATPase with SidK, Rotary State 3
Method: single particle / : Tan YZ

EMDB-11953: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Composite Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-11954: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 2)
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14810: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Consensus Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14811: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Focused Refinement
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-25143: 
Structure of positive allosteric modulator-bound active human calcium-sensing receptor
Method: single particle / : Park J, Zuo H, Frangaj A, Fu Z, Yen LY, Zhang Z, Mosyak L, Slavkovich VN, Liu J, Ray KM, Cao B, Vallese F, Geng Y, Chen S, Grassucci R, Dandey VP, Tan YZ, Eng E, Lee Y, Kloss B, Liu Z, Hendrickson WA, Potter CS, Carragher B, Graziano J, Conigrave AD, Frank J, Clarke OB, Fan QR

EMDB-25144: 
Structure of positive allosteric modulator-free active human calcium-sensing receptor
Method: single particle / : Park J, Zuo H, Frangaj A, Fu Z, Yen LY, Zhang Z, Mosyak L, Slavkovich VN, Liu J, Ray KM, Cao B, Vallese F, Geng Y, Chen S, Grassucci R, Dandey VP, Tan YZ, Eng E, Lee Y, Kloss B, Liu Z, Hendrickson WA, Potter CS, Carragher B, Graziano J, Conigrave AD, Frank J, Clarke OB, Fan QR

EMDB-25145: 
Structure of negative allosteric modulator-bound inactive human calcium-sensing receptor
Method: single particle / : Park J, Zuo H, Frangaj A, Fu Z, Yen LY, Zhang Z, Mosyak L, Slavkovich VN, Liu J, Ray KM, Cao B, Vallese F, Geng Y, Chen S, Grassucci R, Dandey VP, Tan YZ, Eng E, Lee Y, Kloss B, Liu Z, Hendrickson WA, Potter CS, Carragher B, Graziano J, Conigrave AD, Frank J, Clarke OB, Fan QR

EMDB-22738: 
CryoEM reconstruction of SARS-CoV-2 receptor binding domain in complex with the Fab fragment of neutralizing antibody 46
Method: single particle / : Kucharska I, Tan YZ, Benlekbir S, Rubinstein JL, Julien JP

EMDB-22739: 
CryoEM reconstruction of SARS-CoV-2 Spike in complex with the Fab fragment of neutralizing antibody 80.
Method: single particle / : Kucharska I, Tan YZ, Benlekbir S, Rubinstein JL, Julien JP
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
